N. Deepa y M.Y. Sreenivasa
Molecular mycotoxicology Laboratory, Department of Studies in Microbiology, University of Mysore, Manasagangotri, Mysore- 570 006, Karnataka, India.
Correspondence: [email protected]; [email protected]
Feeds are substances or products, including additives, that are either processed, unprocessed or semi-processed and used for feeding animals (Comisión Europea, 2011).
|
They can be classified into of four types:
(EFSA 2016) |
Livestock feed is mainly designed with combinations tailored to the nutritional needs of animals with minimum costs (Pereira et al., 2019).
Cereal and cereal-based products are a major component of animal feed, supplying enormous quantities of nutrients to livestock (Deepthi y Sreenivasa, 2020).

In many developed countries, nearly 70% of harvested cereals are destined to be included in animals’ daily feed, whereas in developing countries they are primarily consumed by humans (Dass et al., 2007).

Globally, feed industries include cereals such as wheat, maize, sorghum, barley, and oats. Among these, wheat and maize are global key commodities in agriculture for animal and poultry feed (FAO, 2009).

A global survey reported that the demand for agricultural commodities is expected to experiment an 84% growth from 2000 to 2050 (FAO y OMS, 2007).
According to Directive 2002/32/EC, poultry and animal feed safety should be prominent, both for governments and producers, since the consumption of feed can pose certain risks to animal health, with hazardous consequences for the human food chain and livestock production industry (FAO 2009; Comisión Europea, 2015).
Due to poor agricultural practices during the pre-harvest, harvest, post-harvest, and storing stages, a high level of mycotoxin-producing fungal contaminants are mainly associated with the cereals that serve as a major ingredient of feed manufacturing (Deepa y Sreenivasa, 2017a, Deepa y Sreenivasa, 2017b) (Figure 1).
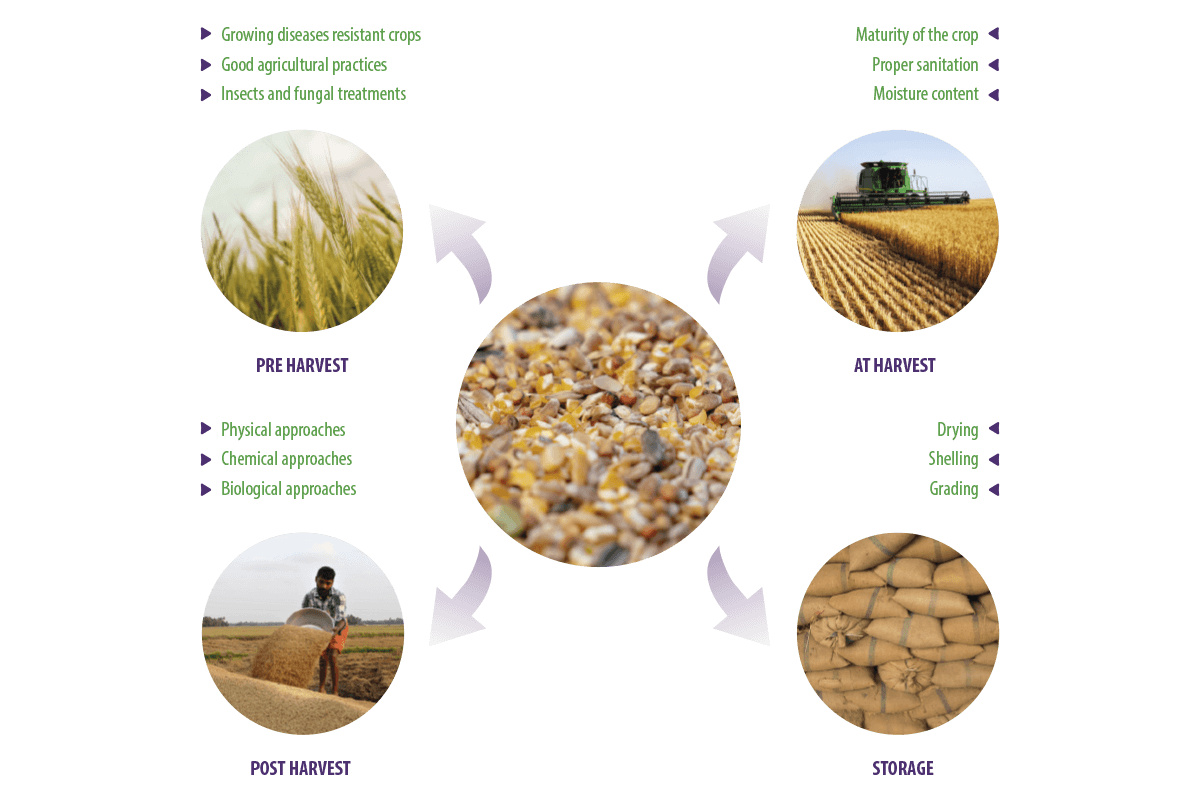
Figure 1. Processing of feed commodities.

The risk of mycotoxins
Mycotoxins are naturally produced by filamentous fungi from the genus Fusarium, Aspergillus, Penicillium, Alternaria, Stachybotrys and Claviceps. Approximately 450 mycotoxins have been reported so far (CAST, 2003).
Based on their occurrence in feed and their consequences on livestock health, certain groups of mycotoxins, such as aflatoxins (AF), fumonisins (FB), ochratoxins (OTA), trichothecenes (TRC), zearalenone (ZEN), and deoxynivalenol (DON) have been targeted in practice through management strategies (Smith et al., 2016).
⇰ Aspergillus species mainly produce AF and OTA, while Penicillium species produce patulin (PAT) and fumonisins, ZEN, DON and trichothecenes are produced by Fusarium species (Iqbal et al., 2015).
The presence of mycotoxigenic fungi in feed has become a significant problem (Kovalsky et al., 2016) as the consumption of the mycotoxins present in the feed has negative health effects ranging from chronic to acute and sometimes even death, as well as affecting animal performance which indirectly affects profitability (CAST, 2003) (Table 1).
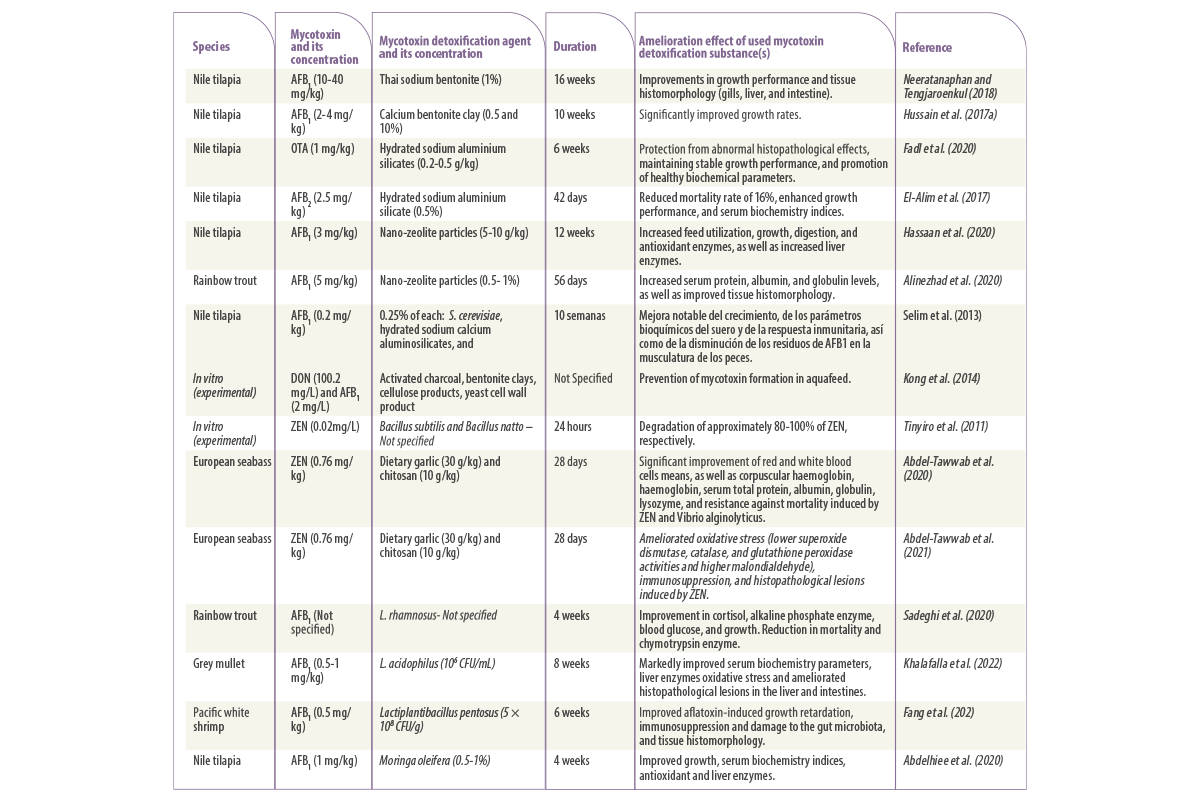
Table 1. Main mycotoxin-producing fungi and their effects on human and animal health.
Worldwide organizations and institutions have restricted the level of mycotoxin acceptance in poultry and animal feeds as mycotoxin-free feed is considered impossible to guarantee (Smith et al., 2016).
Certain standard limits on the level of mycotoxin contamination have been set by legislation, varying in each country since many economic, scientific, and political issues are involved in the decision framing sessions.
For example, the maximum content of AFB1 in feed has been set to 0.02-0.05 ppm and, based on studies, limits for deoxynivalenol, fumonisins, zearalenone and ochratoxin A in feed have been recommended (Comisión Europea 2015; Comisión Europea, 2006).


The importance of mycotoxin detection
The presence of moulds in feed does not directly correlate with the occurrence of its mycotoxin.
Classical culture-based methods used in the detection and identification of mycotoxin producing fungi have various limitations as:
- ⇰ They are labor-intensive, environmentally dependent on the target, and time-consuming.
- ⇰ They have poor reliability of results and difficulties in standardization, among other aspects.
Currently, there is a great demand for molecular technologies to detect mycotoxigenic fungi in feed entering food chain at early stages (Figure 2).
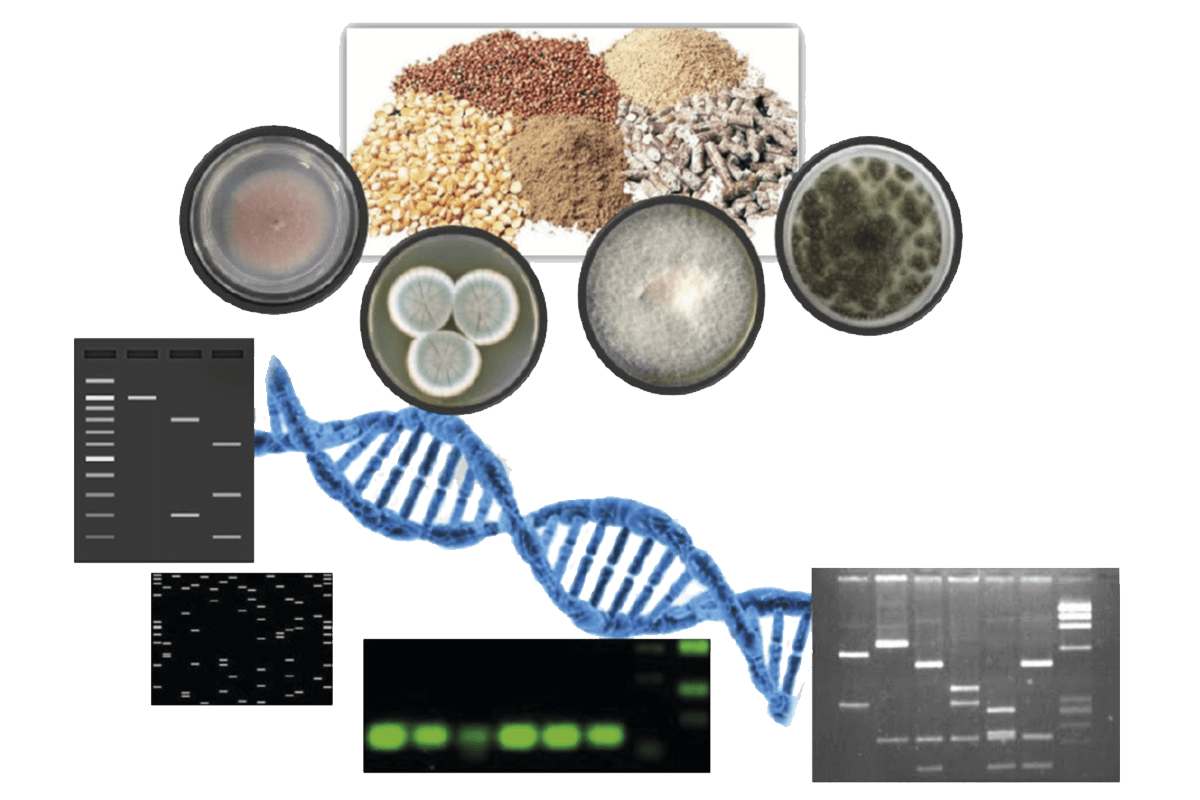
Figure 2. Schematic representation of the detection of fungal contamination in feed and feed matrices through molecular methods.
Researchers have been increasing the use of various molecular techniques in feed industries for the early detection of mycotoxigenic fungi, such as polymerase chain reaction (PCR), nested PCR, quantitative PCR (qPCR), real-time PCR, magnetic capture hybridization PCR (MCH-PCR), immunocapture PCR, in situ PCR, PCR restriction fragment length polymorphism (PCR-RFLP), droplet digilet PCR, cooperational PCR, multiplex PCR, reverse transcriptase PCR (RT-PCR), loop-mediated isothermal amplification (LAMP), DNA microarrays, and fluorescence in situ hybridization (FISH) (Deepa et al., 2021; Deepa et al, 2018) (Figure 3).
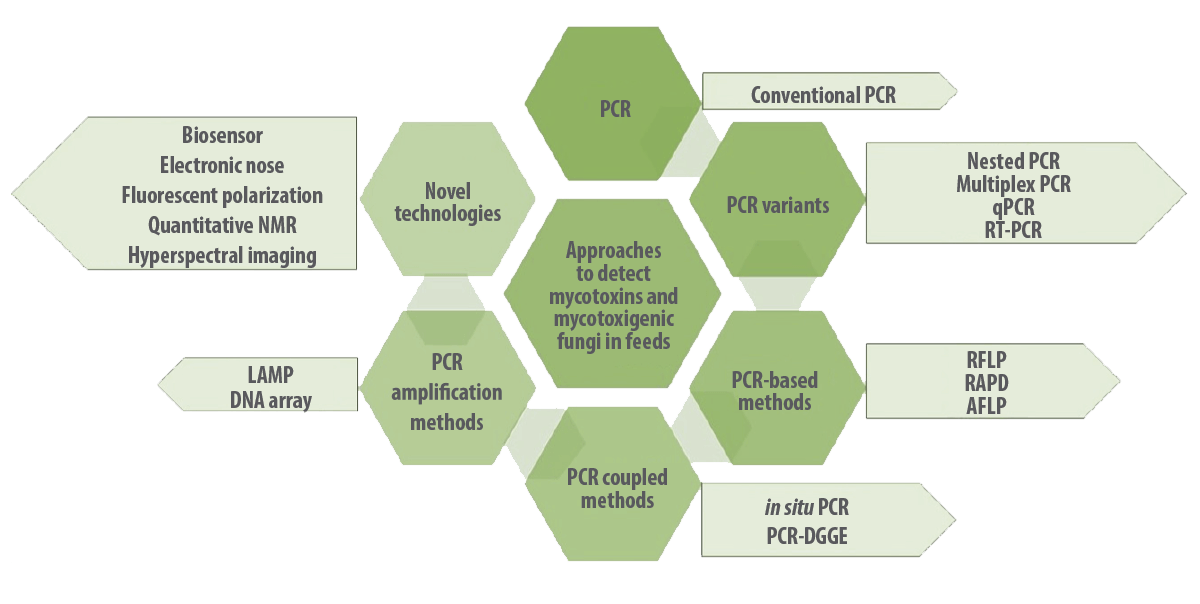
Figure 3. Molecular approaches to detect mycotoxins and mycotoxigenic fungi in feeds and feed matrix.
Different molecular approaches used for the detection and differentiation of mycotoxigenic fungi
Polymerase Chain Reaction (PCR) and its variants
PCR
METHODOLOGY
The PCR technique is used for detecting contamination with mycotoxigenic fungi in feed matrices and to define the potential risk for human and animal health.
⇰ It involves a single round of PCR with a certain set of primers, PCR reagents, and the DNA sample followed by a gel documentation system.
APPLICATION
This technique is mainly applied for the identification of fungi and specific gene detection.
 Fusarium poae producing nivalenol (NIV) was detected through a PCR assay representing the presence of the tri7 gene (Dinolfo et al., 2012).
Fusarium poae producing nivalenol (NIV) was detected through a PCR assay representing the presence of the tri7 gene (Dinolfo et al., 2012). The involvement of F. verticillioides contamination in 51 feed matrices was confirmed through conventional PCR and 69 samples were associated with Fusarium species (Deepa et al., 2016a).
The involvement of F. verticillioides contamination in 51 feed matrices was confirmed through conventional PCR and 69 samples were associated with Fusarium species (Deepa et al., 2016a). Feed material cereal samples were infected with 103 Fusarium isolates and 64 fumonisin-producing F. verticillioides were associated with sorghum (Sreenivasa et al., 2007; 2008).
Feed material cereal samples were infected with 103 Fusarium isolates and 64 fumonisin-producing F. verticillioides were associated with sorghum (Sreenivasa et al., 2007; 2008). Among 108 feed samples, 45.37% of feed mixtures were contaminated with Fusarium species, which was confirmed through a conventional PCR method (Deepthi et al., 2017).
Among 108 feed samples, 45.37% of feed mixtures were contaminated with Fusarium species, which was confirmed through a conventional PCR method (Deepthi et al., 2017).
 Dass et al. (2009) analysed 80 poultry feed mixtures and 114 animal feeds for Fusarium contamination, reporting 374 Fusarium species with primers and 244 isolates for fumonisinspecific strains with FUM1 set of primers.
Dass et al. (2009) analysed 80 poultry feed mixtures and 114 animal feeds for Fusarium contamination, reporting 374 Fusarium species with primers and 244 isolates for fumonisinspecific strains with FUM1 set of primers.

| Advantages | Disadvantages |
| The method is precise and gives rapid results. | It requires much labor and cost-effective technology. |
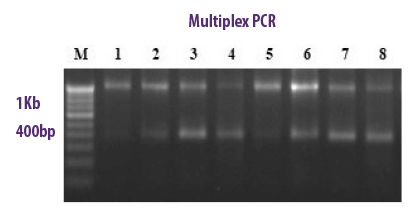
MULTIPLEX PCR
METHODOLOGY
In a single PCR tube, it allows the combination of several species-specific primers used simultaneously to detect a variety of mycotoxigenic fungi. Further amplified products are analysed with gel electrophoresis.
⇰ With this technique a single PCR run is sufficient for the identification of more than one gene at a time.
APPLICATION
 Maize samples associated with F. verticillioides and F. subglutinans contamination were analysed in Brazil by developing gaoB gene coding for galactose oxidase (Faria et al., 2012).
Maize samples associated with F. verticillioides and F. subglutinans contamination were analysed in Brazil by developing gaoB gene coding for galactose oxidase (Faria et al., 2012). One forward and two reverse primers were used to detect fumonisin producing F. verticillioides through mPCR (Deepa et al., 2016c).
One forward and two reverse primers were used to detect fumonisin producing F. verticillioides through mPCR (Deepa et al., 2016c). Aflatoxin-producing and non-aflatoxin-producing Aspergillus species were detected in meju, two sets of primers were used to amplify the genes (omtB, omtA, aflR, ver-1), set I (omtB-1,2, aflR-1,2 and omtA-1,2) and set II (aflR-1,2, ver-1-F-R and omtA-1,2) (Kim et al., 2011). Simultaneously, five major genes, pks (ochratoxin A), pks13 (zearalenone), aflr (aflatoxin), tri5 (trichothecene) and fum13 (fumonisin) produced by Aspergillus, Fusarium and Penicillium were detected using a multiplex PCR assay for feed matrices (Priyanka et al., 2015).
Aflatoxin-producing and non-aflatoxin-producing Aspergillus species were detected in meju, two sets of primers were used to amplify the genes (omtB, omtA, aflR, ver-1), set I (omtB-1,2, aflR-1,2 and omtA-1,2) and set II (aflR-1,2, ver-1-F-R and omtA-1,2) (Kim et al., 2011). Simultaneously, five major genes, pks (ochratoxin A), pks13 (zearalenone), aflr (aflatoxin), tri5 (trichothecene) and fum13 (fumonisin) produced by Aspergillus, Fusarium and Penicillium were detected using a multiplex PCR assay for feed matrices (Priyanka et al., 2015). Four major mycotoxin-related genes, fum13 (fumonisin), nor1 (aflatoxin), Tri6 (trichothecene) and otanps (ochratoxin A) were simultaneously detected in a single test run with specific and sensitive multiplex PCR method (Rashmi et al., 2013).
Four major mycotoxin-related genes, fum13 (fumonisin), nor1 (aflatoxin), Tri6 (trichothecene) and otanps (ochratoxin A) were simultaneously detected in a single test run with specific and sensitive multiplex PCR method (Rashmi et al., 2013).
 Fumonisins (FUM1, FUM13), and trichothecenes (tri5, tri6) were detected in feed matrices using mPCR (Ramana et al., 2011).
Fumonisins (FUM1, FUM13), and trichothecenes (tri5, tri6) were detected in feed matrices using mPCR (Ramana et al., 2011).
| Advantages | Disadvantages |
| The method is accurate, cost effective and a time saver. | Sensitive. |
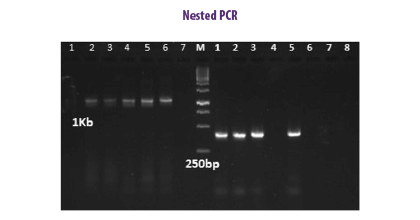
NESTED PCR
METHODOLOGY
This method involves two consecutive reactions in a single closed tube in which the external primer pair amplifies a large amplicon used as template for the internal primer that needs a higher temperature than the external primer and where the internal primer will not bind (Porter-Jordan et al., 1990).
The method is cost-effective and 100 times more sensitive than other methods. Although the risk of contamination may be higher due to the two cycles of amplification, this technique increases the yield and specificity of the target DNA amplification (Rahman et al., 2013).
⇰ The nested PCR method is a double confirmatory test, because if the first primer binds to and amplifies an unwanted DNA sequence, the second set of primers will also bind within the unwanted region.
APPLICATION
 Klemsdal y Elen (2006) detected Fusarium culmorum in cereal crop feed matrices where the external primer pair (FculAF, FculAR) was diluted 10,000 times compared to the internal primer pair (FculBF, FculBR) with a detection limit of 5-50fg of target DNA.
Klemsdal y Elen (2006) detected Fusarium culmorum in cereal crop feed matrices where the external primer pair (FculAF, FculAR) was diluted 10,000 times compared to the internal primer pair (FculBF, FculBR) with a detection limit of 5-50fg of target DNA. Fumonisin producing F. verticillioides associated with feed matrix cereals was detected using one forward (VERF-1) and two reverse primers (VERTR and VERTF-2) (VERTR y VERTF-2) (Deepa et al., 2016b).
Fumonisin producing F. verticillioides associated with feed matrix cereals was detected using one forward (VERF-1) and two reverse primers (VERTR and VERTF-2) (VERTR y VERTF-2) (Deepa et al., 2016b).
| Advantages | Disadvantages |
| Less time-consuming method that avoids frequent DNA isolation for each test run with accurate amplification with less concentration. | Higher risk of contamination. |
REAL-TIME PCR (q-PCR)
METHODOLOGY
This method detects the amplicons of targeted DNA molecules based on the emission of fluorescent signals, which eliminates the post-amplification steps needed in the conventional PCR method, which reduces the time invested. It detects small concentrations and is even useful in identifying in symptomless infections.
APPLICATION
 Even though it can help to detect symptomless infections, qPCR technology is not effective when applied to feed (Ginzinger, 2002).
Even though it can help to detect symptomless infections, qPCR technology is not effective when applied to feed (Ginzinger, 2002).
| Advantages | Disadvantages |
| Time-efficient, sensitive and specific method. | Costly equipment and requirement of costly reagents and expertise to handle the instruments. |
RT-PCR
METHODOLOGY
This technique was developed to differentiate living and dead fungi as mRNA is quickly degraded in dead cells. During the process, mRNA is reverse-transcribed to produce cDNA as a template by a reverse transcriptase enzyme (Brown et al., 2011).
APPLICATION
 F. graminearum was detected using the RT-PCR method in a feed matrix. The method is specific and highly sensitive for detecting mycotoxigenic fungi in feed. However, it is also more expensive and laborious than any other method (Guo et al., 2005).
F. graminearum was detected using the RT-PCR method in a feed matrix. The method is specific and highly sensitive for detecting mycotoxigenic fungi in feed. However, it is also more expensive and laborious than any other method (Guo et al., 2005).
| Advantages | Disadvantages |
| It is a rapid and simple method that does not require agarose gel and allows to obtain qualitative and quantitative data. | The technique requires higher expertise to handle the instruments and it is extremely sensitive with a higher risk of contamination. |
PCR based methods
PCR-RFLP
METHODOLOGY
This method is useful to discriminate at a species level with accuracy, sensitivity, and specificity.
⇰ RFLP records genetic variability among different and same species exhibiting genetic differentiation corresponding with pathogenic and mycotoxicological profiles.
APPLICATION
 Aflatoxin-producing Aspergillus species, such as A. flavus and A. parasiticus, were detected in complex food and feed matrices (Ahmad et al., 2014).
Aflatoxin-producing Aspergillus species, such as A. flavus and A. parasiticus, were detected in complex food and feed matrices (Ahmad et al., 2014). ITS-RFLP discrimination was described between F. verticillioides and F. proliferatum (Visenti et al., 2012).
ITS-RFLP discrimination was described between F. verticillioides and F. proliferatum (Visenti et al., 2012). The PCR-RFLP method was applied to detect variations between 33 fumonisin-producing F. verticillioides strains from feed matrices (Patino et al., 2006).
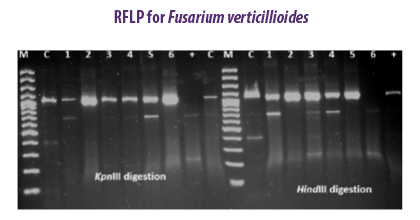
The PCR-RFLP method was applied to detect variations between 33 fumonisin-producing F. verticillioides strains from feed matrices (Patino et al., 2006). Genetic variations were studied among ten strains of F. verticillioides using six different enzymes (EcoRI, Xho I, Hae III, Kpn I, Hinf I, Hind III) isolated from cereal samples (Deepa et al., 2018).
Genetic variations were studied among ten strains of F. verticillioides using six different enzymes (EcoRI, Xho I, Hae III, Kpn I, Hinf I, Hind III) isolated from cereal samples (Deepa et al., 2018).
| Advantages | Disadvantages |
| Simple, reproducible, quick, and stable method that is accurate in the identification of populations. | Requirement of radio isotopes and labor-intensive. |
PCR-AFLP
METHODOLOGY
Amplified fragment Length Polymorphism- PCR is a technique designed to detect DNA polymorphisms.
⇰ Mycotoxicological and fungal pathogen characterization can be complemented by fingerprinting. The method is highly reproducible, sensitive and has the capability to amplify 100 fragments at a time.
APPLICATION
The method was applied to study genetic variability among 3,840 maize kernels in Brazil and FUM1 and FUM8 were the essential genes responsible for fumonisin biosynthesis among F. verticillioides species (Silva et al., 2017).
| Advantages | Disadvantages |
| High reproducibility and the possibility of analyzing different parts of the genome. | Complex and lengthy protocol. |
PCR-RAPD
METHODOLOGY
Random amplified polymorphic DNA identifies genotypic differences using short oligonucleotide primers to distinguish mycotoxigenic fungal species and variations within the species (Zhang et al., 2018).
⇰ Even though the method is used as a diagnostic tool in laboratories, it is not much in practice as it lacks reproducibility (Blackwell et al., 1999).
APPLICATION
 F. subglutinans and F. verticillioides were identified in maize kernels based on RAPD sequences of fragments (Moller et al., 1999).
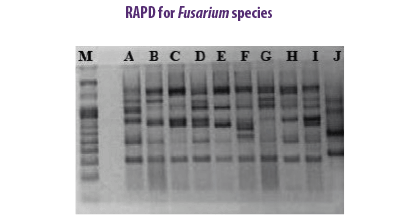
F. subglutinans and F. verticillioides were identified in maize kernels based on RAPD sequences of fragments (Moller et al., 1999). The RAPD-PCR method was applied to ten strains of F. verticillioides associated with cereal samples to study the genetic variation among them (Deepa et al., 2018).
The RAPD-PCR method was applied to ten strains of F. verticillioides associated with cereal samples to study the genetic variation among them (Deepa et al., 2018).
| Advantages | Disadvantages |
| Simple, efficient, and quick method that does not require probes. | Lacking in reproducibility and possible mismatches occur. |
PCR coupled methods
In situ PCR
METHODOLOGY
This technique is a combination of PCR and in situ hybridization to detect mycotoxigenic fungi in tissues or cells (Long, 1998) that has become common for analyzing gene expression and identifying DNA or RNA in cells (Chu et al., 2019). Usually, amplification of DNA from raw materials leads to misinterpretation if the material contains more than one species.
| Advantages | Disadvantages |
| Small tissues can be used at a maximum extent, extensively used due to light microscopy. | Time-consuming method that is difficult to perform with a smaller number of DNA/RNA copies. |
PCR-DGGE
METHODOLOGY
Technique that detects fungal communities by amplifying target DNA, initially using PCR, and later subjected to electrophoresis, but not suitable to describe species present in the sample (Laforgue et al., 2009).
APPLICATION
 Trichoderma harzianum spores from feed matrices were detected through this method. The method is mainly applied in diversity studies and for detecting pathogenic fungi, though it is a time-consuming and poorly reproducible technique (Rytkonen et al., 2011).
Trichoderma harzianum spores from feed matrices were detected through this method. The method is mainly applied in diversity studies and for detecting pathogenic fungi, though it is a time-consuming and poorly reproducible technique (Rytkonen et al., 2011).
| Advantages | Disadvantages |
| Useful in diversity studies. | Lack of reproducibility. |

Post-amplification methods
LAMP
METHODOLOGY
Loop-mediated isothermal amplification is a highly sensitive and successful technique which is directly applied on-site to ensure feed quality and safety, amplifying target DNA in less than an hour at a constant temperature by using four primers to identify six regions on the target DNA with special attention in the annealing step (Tomita et al., 2008).
APPLICATION
 Ochratoxin producing Penicillium nordicum was rapidly detected directly on-site in food and feed production chains (Ferrara et al., 2015).
Ochratoxin producing Penicillium nordicum was rapidly detected directly on-site in food and feed production chains (Ferrara et al., 2015).
| Advantages | Disadvantages |
| Rapid and highly specific method. | High risk of contamination. |
DNA ARRAYS
METHODOLOGY
DNA array technology was developed to detect a large number of organisms at a time. It includes microarray and macroarray techniques in which discrete positioned species-specific detector sequences are fixed on nylon filters or a glass slide. This method is mainly applied in gene expression studies.
⇰ Microarrays consist of a DNA chip for analyzing thousands of mRNAs simultaneously and are mainly used for gene expression studies and applied for the identification of Aspergillus species (Singh y Kumar, 2013).
⇰ Macroarrays involve a DNA hybridization technique and mainly used for identification of A. alternata, F. solani and A. fumigatus in feed matrices (Singh y Kumar, 2013).
APPLICATION
 Kristensen et al. (2007) applied microarray technology for Fusarium species detection which seems to be easy, safe, fast and cost-effective among feed matrices and food. Fusarium species producing 14 trichothecenes and moniliformin detection was determined by DNA microarray.
Kristensen et al. (2007) applied microarray technology for Fusarium species detection which seems to be easy, safe, fast and cost-effective among feed matrices and food. Fusarium species producing 14 trichothecenes and moniliformin detection was determined by DNA microarray.
| Advantages | Disadvantages |
| Quick, easy, bulk analysis that does not require a large amount of DNA sample. | Expensive, time-consuming technique that requires more amount of mRNA. |
Emerging Novel technologies in detection of mycotoxins
Other than Chromatographic methods (Thin Layer Chromatography, Gas Chromatography, and Liquid Chromatography) and Immunoassay-based methods (flow-through membranes, ELISA, dipsticks, Fluorescence Polarization Immunoassay (FPI), and lateral flow immunoassay (LFA)) (Agriopoulou et al., 2020), many more emerging techniques can be applied for detection of mycotoxins in feed.
BIOSENSORS
METHODOLOGY
This fast, easy, reproducible, stable, accurate, and inexpensive on-site-testing method requires transducers (optical, piezoelectric, or electrochemical) for the detection of mycotoxins (Schulz et al., 2019).

APPLICATION
 Oliveria et al. (2019) reported using biosensors for detecting mycotoxins produced by mycotoxigenic fungi in feed and food.
Oliveria et al. (2019) reported using biosensors for detecting mycotoxins produced by mycotoxigenic fungi in feed and food. The method detected AFB1, OTA, AFM1, PAT, ZEN, FB1, T-2, and DON in corn powder, corn, barley, wheat, maize, and corn products (Hossain et al., 2019).
The method detected AFB1, OTA, AFM1, PAT, ZEN, FB1, T-2, and DON in corn powder, corn, barley, wheat, maize, and corn products (Hossain et al., 2019). Impedimetric sensors detected mycotoxigenic fungi producing AFB1, AFM1, OTA, and PAT in food and feedstuffs (Khan et al., 2019).
Impedimetric sensors detected mycotoxigenic fungi producing AFB1, AFM1, OTA, and PAT in food and feedstuffs (Khan et al., 2019). Photentiometric sensors with a two or three-electrode system were used for the successful testing of AFB1 and ZEN in corn powder and maize respectively (He et al., 2019).
Photentiometric sensors with a two or three-electrode system were used for the successful testing of AFB1 and ZEN in corn powder and maize respectively (He et al., 2019). Amperometric sensors based on a two or three-electrode system have also been used to detect ZEN in corn and corn-based products (He et al., 2019).
Amperometric sensors based on a two or three-electrode system have also been used to detect ZEN in corn and corn-based products (He et al., 2019). Surface Plasmon Resonance sensors are sensitive, and cost-effective with high specificity to detect DON, ZEN, and T-2 toxin in wheat (Hossain et al., 2019).
Surface Plasmon Resonance sensors are sensitive, and cost-effective with high specificity to detect DON, ZEN, and T-2 toxin in wheat (Hossain et al., 2019). Quartz crystal microbalance sensors detected AFB1 in rice and wheat (Gu et al., 2019).
Quartz crystal microbalance sensors detected AFB1 in rice and wheat (Gu et al., 2019).
| Advantages | Disadvantages |
| Easy to use, in situ technique with good sensitivity and selectivity. | High cost-effectiveness. |
ELECTRONIC NOSE
METHODOLOGY
E-nose involves non-specific detectors that capture volatile organic compounds (VOCs) and detects qualitative volatile fingerprints associated with toxigenic fungi.
⇰ Odor is the preliminary information to detect metabolite produced on identifying specific VOCs related to fungi on feed matrices.
APPLICATION
 The method detects mycotoxigenic fungi-produced mycotoxins, such as aflatoxins, fumonisins in maize, and DON in wheat bran, which is the major ingredient in feed (Ottoboni et al., 2018).
The method detects mycotoxigenic fungi-produced mycotoxins, such as aflatoxins, fumonisins in maize, and DON in wheat bran, which is the major ingredient in feed (Ottoboni et al., 2018).
| Advantages | Disadvantages |
| Low levels of mycotoxins can be detected on optimization. | The non-volatile nature of mycotoxins is problematic for detection through an E-nose. |
FLUORESCENT POLARIZATION (FP)
METHODOLOGY
FP works with the analyte and the tracer for the antibody binding site. The tracer binding to the antibody influences the tracer molecule’s rotation increasing the FP value.
⇰ The amount of tracer is inversely proportional to the free analyte in the sample, which results in a polarization value inversely proportional to the analyte concentration.
APPLICATION
 The method detects Fusarium species producing ZEN in maize and DON in wheat-based products, as well as Aspergillus species producing AFB1 in maize, and OTA in rice (Huang et al., 2020).
The method detects Fusarium species producing ZEN in maize and DON in wheat-based products, as well as Aspergillus species producing AFB1 in maize, and OTA in rice (Huang et al., 2020). The method is accurate in comparison to HPLC with limited sensitivity (Zhang et al., 2018).
The method is accurate in comparison to HPLC with limited sensitivity (Zhang et al., 2018).
| Advantages | Disadvantages |
| Quick and accurate technique without pre-analytical steps. | Limited sensitivity and expertise are required to handle the instrument. |
QUANTITATIVE NMR AND HYPERSPECTRAL IMAGING
METHODOLOGY
Nuclear magnetic resonance (NMR) spectroscopy is used for the identification of organic compounds and metabolomics. Recently, 2D and multidimensional NMR has been applicable.
Hyperspectral imaging is a technique for the assessment of fungal mycotoxins that works with less cost without destroying the sample, being quick and operating on locating spectral data.
APPLICATION
 NMR-based metabolomics has been used to elucidate the rearrangement mechanism of the Fusarium mycotoxin, Fusarin C, in feed due to its high sample throughput and metabolomics with rich structural information (López-Ruiz et al., 2019).
NMR-based metabolomics has been used to elucidate the rearrangement mechanism of the Fusarium mycotoxin, Fusarin C, in feed due to its high sample throughput and metabolomics with rich structural information (López-Ruiz et al., 2019). Hyperspectral imaging has been mainly used to detect DON and Fusarium pathogen contamination in bulk wheat kernels (Femenias et al., 2020).
Hyperspectral imaging has been mainly used to detect DON and Fusarium pathogen contamination in bulk wheat kernels (Femenias et al., 2020).
| Advantages | Disadvantages |
| Compounds can be elucidated without standards and minimal sample preparation is required. | Lower detection limit, low sensitivity and higher expertise requirements. |

CONCLUSIONS
Certain agricultural commodities, such as wheat, maize, soybeans, bajra, sorghum, and rice, are mainly used for manufacturing animal feed, which they are in higher demand for livestock industries.
Hence, animal feed safety is fundamental as feed consumption is a potential route for hazardous chemicals to enter the human food chain, including mycotoxins.
Due to the threat that mycotoxins pose to animal and human health, certain legislations have been applied in the EU to feed-based products intended for livestock with the establishment of verified limitation criteria, as well as the development of sensitive and consistent methods to detect mycotoxins.
Enormous PCR-based technologies and novel analytical techniques are applied to detect mycotoxins and mycotoxigenic fungi.
Currently, many more molecular methods, such as NASBA, FISH, SAGE, Northern blotting, DNA/RNA probe-based method, PCR-ELISA, MCH-PCR, Next generation PCR, Co-operational PCR, RNA-interference are in practice. However, they have not yet been applied for the detection of mycotoxigenic fungi associated with food and feed matrices.
REFERENCES
1. Agriopoulou, S., Stamatelopoulou, E., Varzakas, T. 2020. Advances in analytical and detection of major mycotoxins in foods. Foods, 9, 518.
2. Ahmad M.M., Ahmad M., Ali A., Hamid R., Javed S., Abdin M.Z., 2014. Detection of Aspergillus flavus and Aspergillus parasiticus from aflatoxin-contaminated peanuts and their differentiation using PCR RFLP. Annals of Microbiology 64: 1597-1605.
3. Bindslev L., Oliver R.P., Johansen, B., 2002. In situ PCR for detection and identification of fungal species. Mycological Research 106: 277-279.
4. Blackwell B.A., Gilliam J.T., Savardm M.E., Miller D., Duvick J.P. 1999. Oxidative deamination of hydrolysed fumonisin B1 (AP1) by cultures of Exophiala spinifera. Natural Toxins. 7: 31–38.
5. Brown N.A., Bass C., Baldwin T.K., Chen H., Massot F., Carion P.W., Urban M., Van De Meene A.M., Hammond-Kosack K.E. 2011. Characterisation of the Fusarium graminearum in wheat and floral interaction. Journal of Pathogens.
6. CAST. 2003. Mycotoxins: Risks in Plant, Animal, and Human Systems; CAST: Ames, IA, USA.
7. Chu Y.H., Hardin, H., Zhang, R., Guo, Z. and Lloyd, R.V., in press. In situ hybridization: introduction to techniques, applications and pitfalls in the performance and interpretation of assays. Seminars in Diagnostic Pathology. 36: 336-341.
8. Dass R.S., Sreenivasa M.Y., Janardhana G.R. 2007. High incidence of Fusarium verticillioides in Animal and Poultry feed mixtures produced in Karnataka, India. Plant Pathology Journal. 6:174-178.
9. Dass R.S., Sreenivasa M.Y., Charith Raj A.P., Janardhana G.R. 2009. PCR-based assay for the rapid detection of fumonisin-producing Fusarium species in maize-based animal and poultry feeds in Karnataka, India. Arhieves of Phytopathology and plant protection. 42:8, 796-804
10. Deepa N., Charith Raj Adkar-Purushothama., Sreenivasa M.Y. 2016b. Nested PCR method for early detection of fumonisin producing Fusarium verticillioides in pure cultures, cereal samples and plant parts. Food Biotechnology 30: 18-29.
11. Deepa N., Charith Raj Adkar-Purushothama., Sreenivasa M.Y. 2016c. Multiplex PCR for the early detection of fumonisin producing Fusarium verticillioides. Food Bioscience. 13: 84-88.
12. Deepa N., Nagaraja H., Sreenivasa, M.Y. 2016a. Prevalence of fumonisin producing Fusarium verticillioides associated with cereals grown in Karnataka (India). Food Science and Human Wellness, 5(3), 156-158.
13. Deepa N., Rakesh S and Sreenivasa M.Y. 2018. “Morphological, pathological and mycotoxicological variations among Fusarium verticillioides isolated from cereals”. 3 biotech. 8:105.
14. Deepa N., Sreenivasa M.Y. 2017b. “Fusarium verticillioides, a globally important pathogen of Agriculture and Livestock: A review”. Journal of Veterinary Medicine and Research. 4(4): 1084.
15. Deepa, N., Charith Raj Adkar-Purushothama, Sreenivasa, M.Y. 2021. Chapter 6: “Molecular technologies for the early detection of fungal phytopathogens associated with cereal crops”. Microbial genomic technologies to combat the problems of farming lands. Microbes in land use change management. Elsevier. 137-154.
16. Deepa, N., Sreenivasa, M.Y. 2017a. “Fumonisins: A review on its Global Occurrence, Epidemiology, Toxicity and Detection”. Journal of Veterinary Medicine and Research. 4(6): 1093.
17. Deepthi, B.V., Gnanaprakash, A.P., Sreenivasa M.Y. 2017. Effet of γ irradiation on fumonisin producing Fusarium associated with animal and poultry feed mixtures. 3 Biotech. 7:57.
18. Deepthi, B.V., Sreenivasa M.Y. 2020. Fumonisins – The underrated mycotoxins in poultry, livestock and humans. mycotoxinsite. Com
19. Dinolfo, M., Barros, G. and Stenglein, S., 2012. Development of a PCR assay to detect the potential production of nivalenol in Fusarium poae. FEMS Microbiology Letters 332: 99-104.
20. EU Commission. 2015. The European Parliament and The Council of the European Union Directive 2002/32/EC of 22 the European Parliament and of the Council of 7 May 2002 on undesirable substances in animal feed. Official Journal of the European Union. 32, 1–30.
21. European Commission. Comission recomendation of 14 January 2011 establishing guidelines for the distinction between feed materials, feed additives, biocidal products and veterinary medicinal products. 2011. Official Journal of the European Union. 75–79.
22. FAO & WHO. 2007. Animal Feed Impact on Food Safety; FAO: Rome, Itlay, 23. FAO. 2009. The State of Food and Agriculture; FAO: Rome, Italy. ISBN 9789251062159.
24. Faria C.B., Abe C.A.L., Novais da- silva C., Tessmann D.J., Barbosa Tesmann L.P. 2012. New PCR assays for the identification of Fusarium verticillioides, F. subglutinans, and other species of the Gibberella fujikuroi complex. International Journal of Molecular Sciences. 13: 115-132.
25. Femenias, A., Gatius, F., Ramos, A.J., Sanchis, V., Marín, S. 2020. Use of hyperspectral imaging as a tool for Fusarium and deoxynivalenol risk management in cereals: A review. Food Control. 108, 106819.
26. Ferrara M., Perrone G., Gallo A., Epifani F., Visconti A., Susca A., 2015. Development of loop-mediated isothermal amplification (LAMP) assay for the rapid detection of Penicillium nordicum in dry- cured meat products. International Journal of Food Microbiology. 202: 42-47.
27. Food Standards Agency Food.gov.uk. Available online: https://www.food.gov.uk/business-industry/farmingfood/animalfeed/what-farmanimals-eat (accessed on 4 December 2016).
28. Ginzinger D.G. 2002. Gene quantification using real-time quantitative PCR: An emerging technology hits the mainstream. Experimental Hematology. 30: 503-512.
29. Gu, Y., Wang, Y., Wu, X., Pan, M., Hu, N., Wang, J., Wang, S. 2019. Quartz crystal microbalance sensor based on covalent organic framework composite and molecularly imprinted polymer of poly(o-aminothiophenol) with gold nanoparticles for the determination of aflatoxin B1. Sensors and Actuators B Chemistry. 291, 293–297.
30. Guo J.R., Schnieder F., Beyer M., Verreet, J.A., 2005. Rapid detection of Mycosphaerella graminicola in wheat using reverse transcription- PCR assay. Journal of Phytopathology 153: 674-679.
31. He, B., Yan, X. 2019. An amperometric zearalenone aptasensor based on signal amplification by using a composite prepared from porous platinum nanotubes, gold nanoparticles and thionine-labelled graphene oxide. Microchimica Acta 2019, 186, 383.
32. Hossain, Z., Busman, M., Maragos, C.M. 2019. Immunoassay utilizing imaging surface plasmon resonance for the detection of cyclopiazonic acid (CPA) in maize and cheese. Analytical and Bioanalytical Chemistry. 411, 3543–3552.
33. Huang, X., Tang, X., Jallow, A., Qi, X., Zhang, W., Jiang, J., Li, H., Zhang, Q., Li, P. 2020. Development of an Ultrasensitive and Rapid Fluorescence Polarization Immunoassay for Ochratoxin A in Rice. Toxins. 12, 682.
34. Khan, R., Ben Aissa, S., Sherazi, T.A., Catanante, G., Hayat, A., Marty, J.L. 2019. Development of an Impedimetric Aptasensor for Label Free Detection of Patulin in Apple Juice. Molecules. 24, 1017.
35. Kim S.W., Li Z., Moore P.S., Monaghan A.P., Chang Y., Nichols M., John B. 2010. A sensitive non-radioactive northern blot method to detect small RNAs. Nucleic Acids Research. 38: 98-110.
36. Klemsdal S.S., Elen O. 2006. Development of a highly sensitive nested PCR method using a single closed tube for detection of Fusarium culmorum in cereal samples. Letters in Applied microbiology. 42: 544-548.
37. Kristensen R., Gauthier G., Berdal K.G., Hamels S., Remacle J., Holst Jensen A., 2007. DNA microarray to detect and identify trichotheceneand moniliformin-producing Fusarium species. Journal of Applied Microbiology 102: 1060-1070.
38. Laforgue R., Guérin L., Pernelle J.J., Monnet C., Dupont J., Bouix M., 2009. Evaluation of PCR-DGGE methodology to monitor fungal communities on grapes. Journal of Applied Microbiology 107: 1208-1218.
39. Long A.A. 1998. In situ polymerase chain reaction: foundation of the technology and today’s options. European Journal of Histochemistry 42: 101-109.
40. López-Ruiz, R., Romero-González, R., Frenich, A.G. 2019. Metabolomics approaches for the determination of multiple contaminants in food. Current Opinion in Food Science. 28, 49–57.
41. Moeller E.M., Chelkowski J., Geiger H.H. 1999. Species-specific PCR assays for the fungal pathogens Fusarium moniliforme and Fusarium subglutinans and their application to diagnose maize ear rot. Journal of Phytopathology. 147: 497-508.
42. Oliveira, I.S., Junior, A.G.D.S., De Andrade, C.A.S., Oliveira, M.D.L. 2019. Biosensors for early detection of fungi spoilage and toxigenic and mycotoxins in food. Current Opinion in Food Science. 29, 64–79.
43. Ottoboni, M., Pinotti, L., Tretola, M., Giromini, C., Fusi, E., Rebucci, R., Grillo, M., Tassoni, L., Foresta, S., Gastaldello, S. et al. 2018. Combining E-Nose and Lateral Flow Immunoassays (LFIAs) for Rapid Occurrence/Co-Occurrence Aflatoxin and Fumonisin Detection in Maize. Toxins. 10, 416.
44. Pereira, C.S., Cunha S.C., Fernandes, J.O. 2019. Prevalent mycotoxins in animal feed: occurrence and analytical methods. Toxins. 11, 290.
45. Porter-Jordan, K., Rosenberg, E.I., Keiser, J.F., Gross, J.D., Ross, A.M., Nasim, S. and Garrett, C.T., 1990. Nested polymerase chain reaction assay for the detection of cytomegalovirus overcomes false positives caused by contamination with fragmented DNA. Journal of Medical Virology 30: 85-91.
46. Priyanka S.R., Mudili D.V., Balakrishna K.S., Murali H., Batra H., 2015. Development and evaluation of a multiplex PCR assay for simultaneous detection of major mycotoxigenic fungi from cereals. Journal of Food Science and Technology 52: 486-492.
47. Rahman H, Ur., Yue X., Yu Q., Zhang W., Zhang Q., Li P. 2020. Current PCR-based methods for the detection of mycotoxigenic fungi in complex food and feed matrices. World Mycotoxin Journal, 13 (2): 139-150.
48. Ramana M.V., Balakrishna K., Murali H.C.S., Batra H.V. 2011. Multiplex PCR based strategy to detect contamination with mycotoxigenic Fusarium species in rice and finger millet collected from southern India. Journal of the Science of Food and Agriculture. 91: 1666-1673.
49. Rashmi R., Ramana M.V., Shylaja R., Uppalapati S.R., Murali H.S., Batra H.V., 2013. Evaluation of a multiplex PCR assay for concurrent detection of four major mycotoxigenic fungi from foods. Journal of Applied Microbiology 114: 819-827.
50. Rytkönen K.T., Williams T.A., Renshaw G.M., Primmer C.R., Nikinmaa M. 2011. Molecular evolution of the metazoan PHD-HIF oxygen-sensing system. Molecular Biology and Evolution. 28: 1913–1926
51. Schulz, K., Pöhlmann, C., Dietrich, R., Märtlbauer, E., Elßner, T. 2019. Electrochemical Biochip Assays Based on Anti-idiotypic Antibodies for Rapid and Automated On-Site Detection of Low Molecular Weight Toxins. Frontiers in Chemistry. 7, 31.
52. Silva J.J., Viaro H.P., Ferranti L.S., Oliveira A.L.M., Ferreira J.M., Ruas C.F., Ono E.Y.S., Fungaro M.H.P. 2017. Genetic structure of Fusarium verticillioides populations and occurrence of fumonisins in maize grown in southern Brazil. Crop Protection. 99: 160-167.
53. Singh A., Kumar N., 2013. A review on DNA microarray technology. International Journal of Current Research and Reviews. 5: 01–05.
54. Smith M.C., Madec S., Coton E., Hymery N. 2016. Natural Co-occurrence of mycotoxins in foods and feeds and their in vitro combined toxicological effects. Toxins. 8, 94.
55. Sreenivasa M.Y., Dass R.S., Charithraj A.P., Janardhana G.R., 2007. PCR method for the detection of genus Fusarium and fuonisin producing isolates from freshly harvested sorghum grains grown in Karnataka, India. J food saf. 28, 236-247.
56. Sreenivasa M.Y., Gonzalez Jaen M.T., Dass R.S., Charithraj A.P., Janrdhana G.R., 2008. A PCR based assay for the detetction and differentiation of potential fumonisin producing Fusarium verticillioides isolated from maize kernels. Food biotechnol. 22, 160-170.
57. Tomita N., Mori Y., Kanda H., Notomi T., 2008. Loop-mediated isothermal amplification (LAMP) of gene sequences and simple visual detection of products. Nature Protocols 3: 877-882.
58. Visentin I., Montis V., Doll K., Alabouvette C., Tamietti G., Karlovsky P., Cardinale F. 2012. Transcription of genes in biosynthetic pathway for fumonisin mycotoxins is epigenetically and differentially regulated in the fungal maize pathogen Fusarium verticillioides. Eukaryotic cell. 11: 252-259.
59. Zhang, X., Tang, Q., Mi, T., Zhao, S., Wen, K., Guo, L., Mi, J., Zhang, S., Shi, W., Shen, J., et al. 2018. Dual-wavelength fluorescenc polarization immunoassay to increase information content per screen: Applications for simultaneous detection of total aflatoxin and family zearalenones in maize. Food Control. 87, 100–108.
60. Zhao Y., Liu N., Niu, J.X. 2009. A study of the distribution of apple stem pitting virus in tissues of pear tree using in situ hybridization and in situ RT-PCR. Agricultural Sciences in China 8: 1351-1359.




 Micotoxicosis prevention
Micotoxicosis prevention